| Outline CV | My publications arranged by subject; with links to pdf files |
|
| Presentations used in teaching and research:These are PowerPoint presentations and all include links to download the original presentations if required. These were prepared for use in the University of East Anglia and Sri Venkateswara University in Tirupati, India. Topics include Introduction to Microbial metabolism, Bioenergetics, Methylotrophy | ||
| Brief summary of my research career with list and pictures of my research students | ||
| Full accounts of my research | ||
| Websites I use [to data bases, literature, protein structure manipulations] | ||
| The Quinoprotein Symposium I organisised in 2002 in Southampton | ||
| Bits and pieces (for my use) | ||
The Biochemistry of Methylotrophs. Microbes that are able to grow on compounds with only one carbon atom [C1-compounds] such as methane, methanol, methylamine etc are called Methylotrophs. Those that grow on methane are also called Methanotrophs.All the earlier work on these microbes [up to 1981] is described in
Anthony (1982): The Biochemistry of Methylotrophs. |
||||||||||||||||||||||||||||||||||||||||||||||
| TOP | ||||||||||||||||||||||||||||||||||||||||||||||
The PQQ-containing quinoproteins. These are the equivalent of flavoproteins that have the riboflavin derivatives FMN or FAD as their prosthetic groups. |
||||||||||||||||||||||||||||||||||||||||||||||
.jpg) .jpg) |
||||||||||||||||||||||||||||||||||||||||||||||
| The 'propeller' structure of MDH | ||||||||||||||||||||||||||||||||||||||||||||||
 |
||||||||||||||||||||||||||||||||||||||||||||||
| The 'Tryptophan docking' system that holds together the propeller blades | ||||||||||||||||||||||||||||||||||||||||||||||
.jpg) .jpg) |
||||||||||||||||||||||||||||||||||||||||||||||
.jpg) .jpg) |
||||||||||||||||||||||||||||||||||||||||||||||
| TOP OF QUINOPROTEINS TOP OF MY RESEARCH |
||||||||||||||||||||||||||||||||||||||||||||||
| The electron transport chain from MDH to oxygen: This is very unusual as it consists of 3 periplasmic proteins and a typical cytochrome oxidase. The primary electron acceptor from MDH is a completely novel type of cytochrome called cytochrome cL; this passes electrons to a more typical small cytochrome cH which is the electron donor to the oxidase. | ||||||||||||||||||||||||||||||||||||||||||||||
.jpg) |
||||||||||||||||||||||||||||||||||||||||||||||
| Cytochrome cL: The structure of cytochrome cL from
Methylobacterium extorquens has been
determined by X-ray crystallography
to a resolution of 1.6 A ° . This
unusually large, acidic cytochrome is the physiological electron acceptor
for the quinoprotein methanol dehydrogenase in the periplasm of
methylotrophic bacteria. Its amino acid sequence is completely different
from that of other cytochromes but its X-ray structure reveals a core that is
typical of class I cytochromes c, having a-helices folded into a compact
structure enclosing the single haem c prosthetic group and leaving one
edge of the haem exposed. The haem is bound through thioether bonds to
Cys65 and Cys68, and the fifth ligand to the haem iron is provided by
His69. Remarkably, the sixth ligand is provided by His112, and not by
Met109, which had been shown to be the sixth ligand in solution. Cytochrome cL is unusual in having a disulphide bridge that tethers the long C-terminal extension to the body of the structure. The crystal structure reveals that, close to the inner haem propionate, there is tightly bound calcium ion that is likely to be involved in stabilization of the redox potential, and that may be important in the flow of electrons from reduced pyrroloquinoline quinone in methanol dehydrogenase to the haem of cytochrome cL. As predicted, both haem propionates are exposed to solvent, accounting for the unusual influence of pH on the redox potential of this cytochrome. For Protein Data Bank file [pdb]: cytochrome cL |
||||||||||||||||||||||||||||||||||||||||||||||
  (2).jpg) |
||||||||||||||||||||||||||||||||||||||||||||||
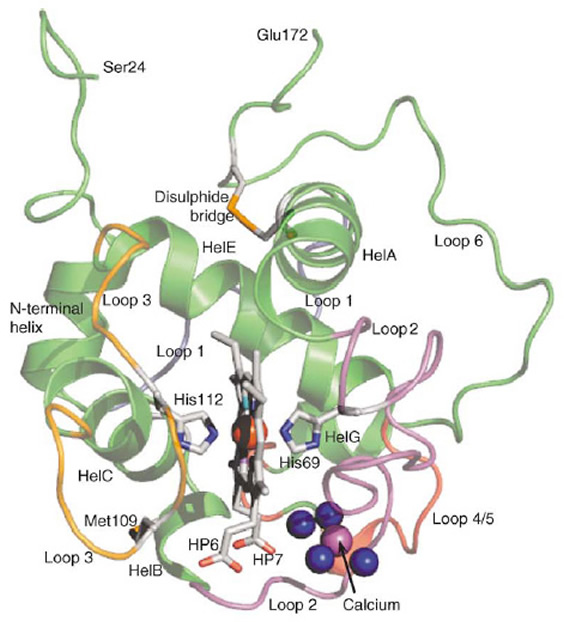
| The main structural features of cytochrome cL. Although there is no sequence identity with other cytochromes, the helices A, C and E constitute the typical haem-enclosing fold seen in all cytochromes c. Helices are labelled HelA, HelB, etc. Loop 1 (grey) joins the N-terminal helix and helix A; loop 2 (purple) between helix A and helix B, carries the haem-binding sequence and the amino acid residues that coordinate to the calcium ion; loop 3 (orange) is the exceptionally flexible loop that joins helix C to helix E, and carries the sixth ligand to the haem (His112) and the methionine (Met109) that is the sixth ligand in solution. The red sphere is the iron atom at the centre of the haem prosthetic group. HP6 is the outer haem propionate group and HP7 is the inner haem propionate group. The blue spheres are the water molecules (Wat6-Wat9) that coordinate to the calcium ion (magenta sphere). Met109 is the residue that forms the sixth ligand to the haem in solutions of the cytochrome. | ||||||||||||||||||||||||||||||||||||||||||||||
| TOP | ||||||||||||||||||||||||||||||||||||||||||||||
| Cytochrome cH: This small cytochrome mediates electron flow from cytochrome cL to the oxidase. Cytochrome cH is the electron donor to the oxidase in methylotrophic bacteria. Its amino acid sequence suggests that
it is a typical Class I cytochrome c, but some features of the sequence indicated that its structure might be of special
interest. The structure of oxidized cytochrome cH has been solved to 2.0 Å resolution by X-ray diffraction. It has the
classical tertiary structure of the Class 1 cytochromes c but bears a closer gross resemblance to mitochondrial cytochrome
c than to the bacterial cytochrome c2. The left-hand side of the haem cleft is unique; in particular, it is highly
hydrophobic, the usual water is absent, and the “conserved” Tyr67 is replaced by tryptophan. A number of features of
the structure demonstrate that the usual hydrogen bonding network involving water in the haem channel is not essential
and that other mechanisms may exist for modulation of redox potentials in this cytochrome. For Protein Data Bank file [pdb]: Cytochrome cH |
||||||||||||||||||||||||||||||||||||||||||||||
.jpg) .jpg) |
||||||||||||||||||||||||||||||||||||||||||||||
TOP OF MY RESEARCH
TOP OF QUINOPROTEINS
|
||||||||||||||||||||||||||||||||||||||||||||||
My presentation to a conference on vitamins in Osaka
and to Mitsubishi scientists in Niigata
Click here for my photos of this visit
| Introduction: PQQ cannot seriously be accepted as a vitamin until an enzyme can be shown to require it as its cofactor. About one year ago Kasahara and Kato claim to have provided this evidence and announced ‘A new redox-cofactor vitamin for mammals' in Nature. This was greeted with enthusiasm by Reuters news agency “The first new vitamin for 55 years”, and its exploitation by Mitsubishi seems to be underway (put vitamin and PQQ into Google for a good survey of this). However, the claim of Kasahara and Kato was based on sequence analysis of an enzyme, predicted to be involved in mouse lysine metabolism, using databases and search engines which inappropriately label beta propeller sequences as PQQ-binding sites. The ‘sites' wrongly identified by the databases do not represent PQQ-binding sites but represent the Beta -sheets that form the ‘blades' of the ‘propeller fold' which happens to be a feature of all PQQ-dependent dehydrogenases, whose main structure is a superbarrel made up of either six or eight ‘propeller blades'. What the evidence actually suggests is that their (predicted) enzyme is an interesting novel protein having an eight-bladed beta propeller structure; but there is no evidence that it is a PQQ-dependent dehydrogenase. There is also no evidence that this protein has any relevance to lysine metabolism. This material has been published the journal Nature (Felton and Anthony, 2005) (Download PDF). |
Dr Leigh M. Felton |
 |
.jpg) |
|
.jpg) |
|
.jpg) |
|
.jpg) |
|
.jpg) |
|
.jpg) |
|
.jpg) |
|
.jpg) |
|
.jpg) |
|
.jpg) |
|
.jpg) |
|
.jpg) |
|
.jpg) |
|
.jpg) |
|
TOP of page

.jpg)




.jpg)
.jpg)
.jpg)
